Laboratory of Systems Tumor Immunology Erlangen, Germany
Menu
January 2022. We initiate a project on bioinformatics ans computer-model based design of CART cells funded by the Hiege Stiftung.
October 2021. We got a paper published about deep learning and networks applied to melanoma at the prestigious "International Journal of Cancer". See the paper here.
January 2021. In our IJMS paper we collect ideas and strategies to convince clinicians and experimentalists about the advantages of mathematical modelling. see the paper here
November 2020. Our work in engineering of dendritic cells with miRNAs for immunotherapy. see the paper here
February 2019. Our work in network biology applied to oligodendrocyte differentiation is out. see the paper here
January 2019. The book we edited in ncRNAs is out. Check it here
November 2018. The paper with our collaborators in Charité Berlin on antibitics and lung infection is published. See here
April 2 2018. Our web platform for network-based mining of HTD in melanoma is published
January 1 2018. In this new paper we explore the use of cooperative miRNAs to fight cancer resistance
August 1 2017. Our work on E2F1-centered EMT networks in cancer accepted in Nature Communications!
July 1 2017. Beautiful work combining network analysis and modelling to understand brain cell differentiation.
21 February 2017. Our paper on identification of miRNA-mRNA network modules in colon cancer is out
30 January 2017. Our paper on web platform for network analysis of macrophage HTD is out
20 December 2016. The Staedler Foundation is to support our work on uveal melanoma networks
26 July 2016. Our review on mathematical modelling of miRNA regulation published in Nucleic Acids Research
27 April 2016. Our paper with the Schmeck group in Marburg on microRNA-feedback loops in Streptococcus pneumoniae lung infection is published!
October 2021. We got a paper published about deep learning and networks applied to melanoma at the prestigious "International Journal of Cancer". See the paper here.
January 2021. In our IJMS paper we collect ideas and strategies to convince clinicians and experimentalists about the advantages of mathematical modelling. see the paper here
November 2020. Our work in engineering of dendritic cells with miRNAs for immunotherapy. see the paper here
February 2019. Our work in network biology applied to oligodendrocyte differentiation is out. see the paper here
January 2019. The book we edited in ncRNAs is out. Check it here
November 2018. The paper with our collaborators in Charité Berlin on antibitics and lung infection is published. See here
April 2 2018. Our web platform for network-based mining of HTD in melanoma is published
January 1 2018. In this new paper we explore the use of cooperative miRNAs to fight cancer resistance
August 1 2017. Our work on E2F1-centered EMT networks in cancer accepted in Nature Communications!
July 1 2017. Beautiful work combining network analysis and modelling to understand brain cell differentiation.
21 February 2017. Our paper on identification of miRNA-mRNA network modules in colon cancer is out
30 January 2017. Our paper on web platform for network analysis of macrophage HTD is out
20 December 2016. The Staedler Foundation is to support our work on uveal melanoma networks
26 July 2016. Our review on mathematical modelling of miRNA regulation published in Nucleic Acids Research
27 April 2016. Our paper with the Schmeck group in Marburg on microRNA-feedback loops in Streptococcus pneumoniae lung infection is published!
26th May 2015. We just published our paper on rough hypercuboid clustering of miRNAs!
May 2015. Our group member Pia Wentker has successfully defended her PhD Thesis. Congratulations, Dr. Wentker!
1st March 2015. Our paper on G2-M arrest in melanoma cells controlled by the interaction between Chk1 and Wee1 is accepted!
|
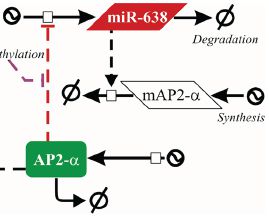
19th January 2015. Our paper on miR-638 and melanoma metastasis is accepted!
|
|
12th December 2014. Our review entitled "Envisioning the Application of Systems Biology in Cancer Immunology" is now published by Springer.
|
|
1st September 2014. Our review in Cancer Systems Biology published in Wiley´s eLS Citable Reviews in the Life Sciences.
|
|
24th July 2014. Dr. Lai selected to participate in the Computational Biology Workshop organized by Bayer in Boston USA, September 13th – 16th, 2014. He will present our results in miRNA regulation
|
21st July 2014. We have been selected to participate in the SPP 1757 Priority
Program of the Deutsche Forschungsgemeinschaft (DFG). We will woprk together with Prof. Wegner dissecting the networks modulating the diffrentiation of oligodendrocytes.
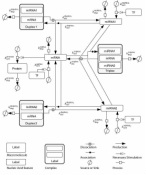
2nd July 2014. Our model in miRNA cooperativity has beed selected "Model of the Month Article" in BioModels Database

18th June 2014. Our work in miRNA cooperativity presented in major Systems Medicine Conference. Our collaborator from the University of Rostock Ulf Schmitz is presenting our work in cooperative miRNAs in the International Conference on Systems Biology of Human Disease 2014, Boston, USA.
http://www.sbhd2014.org/
http://www.sbhd2014.org/

17th June 2014. The book on Cancer miRNA edited by Prof. Vera together with with former colleagues from Rostock ULf Schmitz and Olaf Wolkenhauer has been a success during his first year. Chapters from our book have been downloaded more that 27,000 times and that it is among the 25% best performing books. Some chapters of the book are cited more than 150 times according to Google Scholar.
You can find the book here.
And our book chapter on System Biology of miRNAs here.
You can find the book here.
And our book chapter on System Biology of miRNAs here.
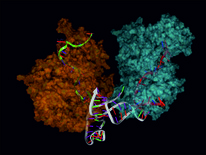
10th June 2014. Our paper on genome wide analysis of miRNA cooperativity is published.
Cooperative gene regulation by microRNA pairs and their identification using a computational workflow. Nucl. Acids Res. (2014) doi: 10.1093/nar/gku465
MicroRNAs (miRNAs) are an integral part of gene regulation at the post-transcriptional level. Recently, it has been shown that pairs of miRNAs can repress the translation of a target mRNA in a cooperative manner, which leads to an enhanced effectiveness and specificity in target repression. However, it remains unclear which miRNA pairs can synergize and which genes are target of cooperative miRNA regulation. In this paper, we present a computational workflow for the prediction and analysis of cooperating miRNAs and their mutual target genes, which we refer to as RNA triplexes. The workflow integrates methods of miRNA target prediction; triplex structure analysis; molecular dynamics simulations and mathematical modeling for a reliable prediction of functional RNA triplexes and target repression efficiency. In a case study we analyzed the human genome and identified several thousand targets of cooperative gene regulation. Our results suggest that miRNA cooperativity is a frequent mechanism for an enhanced target repression by pairs of miRNAs facilitating distinctive and fine-tuned target gene expression patterns. Human RNA triplexes predicted and characterized in this study are organized in a web resource at www.sbi.uni-rostock.de/triplexrna/.
Cooperative gene regulation by microRNA pairs and their identification using a computational workflow
Cooperative gene regulation by microRNA pairs and their identification using a computational workflow. Nucl. Acids Res. (2014) doi: 10.1093/nar/gku465
MicroRNAs (miRNAs) are an integral part of gene regulation at the post-transcriptional level. Recently, it has been shown that pairs of miRNAs can repress the translation of a target mRNA in a cooperative manner, which leads to an enhanced effectiveness and specificity in target repression. However, it remains unclear which miRNA pairs can synergize and which genes are target of cooperative miRNA regulation. In this paper, we present a computational workflow for the prediction and analysis of cooperating miRNAs and their mutual target genes, which we refer to as RNA triplexes. The workflow integrates methods of miRNA target prediction; triplex structure analysis; molecular dynamics simulations and mathematical modeling for a reliable prediction of functional RNA triplexes and target repression efficiency. In a case study we analyzed the human genome and identified several thousand targets of cooperative gene regulation. Our results suggest that miRNA cooperativity is a frequent mechanism for an enhanced target repression by pairs of miRNAs facilitating distinctive and fine-tuned target gene expression patterns. Human RNA triplexes predicted and characterized in this study are organized in a web resource at www.sbi.uni-rostock.de/triplexrna/.
Cooperative gene regulation by microRNA pairs and their identification using a computational workflow